 In addition, users can provide their own networks and nomenclature datasets. StrongestPath comes with two types of built-in databases: (I) some PPI and signaling networks of human and mouse, containing interactions recorded in public databases, (II) protein nomenclature database, containing 11 different symbols and accession IDs of genes and proteins in different databases. For example, when a list of genes is identified in a study of a phenomenon, researchers seek to answer whether there is a regulatory pathway between the transcription factors associated with the phenomenon and the identified genes regarding the experimentally validated data reported in the public databases. The third challenge is identifying any activating or inhibitory regulatory path between two distinct groups of proteins. This feature can be used to identify unknown elements of a protein complex, biological process or core regulatory circuitry. To address this challenge, StrongestPath looks at the whole PPI or signaling network and identifies proteins with maximum total confidence of interactions with the given set of proteins. The second challenge addressed by StrongestPath is growing the sub-network of the input proteins, either by extracting their pairwise interactions from a list of PPI or signaling databases, or adding further proteins that are more likely to create protein complexes or dense interactions with the input proteins. In many experimental studies perturbation of a protein A is observed to influence a protein B, but the cascade of interactions between A and B is unidentified. The first challenge is identifying a cascade of interactions, as a regulatory or signaling pathway, in a large PPI or signaling network. We developed StrongestPath, a Cytoscape 3.0 App, to address three key challenges during analysis of PPI or signaling networks. PesCa, PathExplorer ( ) and PathLinker, are examples of such apps that can compute paths in biological networks. Other Plugins and Apps can be integrated into this flexible platform for complex network analysis and visualisations. To make this process easier, Cytoscape was developed to help with molecular and network profiling analysis and also visualizing molecular interaction networks. So, the identification of the cascades of interactions from the receptors to the transcriptional regulatory factors is a major challenge in systems biology. higher confidences for the experimentally validated interactions, and lower values for the computationally predicted ones. Some of the databases assign different confidence levels to the interactions, e.g. 
There have been numerous protein–protein interaction (PPI) or signaling pathways databases developed based on the experimental approaches or computational predictions.  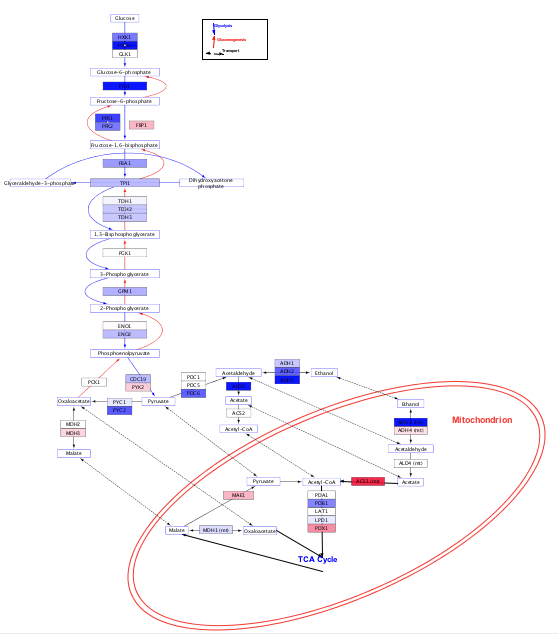
The teamwork of proteins, in terms of temporary or permanent interactions, is critical for any biological process. The Creative Commons Public Domain Dedication waiver ( ) applies to the data made available in this article, unless otherwise stated in a credit line to the data. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. Open AccessThis article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
